Semantic Representation of Scientific Literature (PeerJ CompSci 2015): Supplementary Material
This page documents our supplementary material for the PeerJ submission [1]: "Semantic representation of scientific literature: bringing claims, contributions and named entities onto the Linked Open Data cloud", PeerJ Computer Science, vol. 1, no. e37 PeerJ, 12/2015.
1. Data
1.1. Corpora
In our paper, we intermittently refer to the documents in our experiments in the Design, Evaluation and Application sections. The three open-access corpora used in our experiments are as follows:
- The proceedings of the Semantic Publishing (SePublica) workshop from 2011–2014.
- Open-access papers from the computer science edition of PeerJ journal.
- Argumentative Zoning (AZ) corpus, which is a collection of 80 conference articles in computational linguistics, originally curated by Simone Teufel.
To resolve the documents IDs mentioned in the paper to their actual URLs, please refer to the corpus-index file.
1.2. Gold Standard
For the intrinsic evaluation of our text mining pipeline for RE extraction, we manually curated a gold standard corpus of 30 documents from three different corpora described above. Each sentence in a document that contained a rhetorical entity was manually annotated and classified as either a Claim or Contribution by adding the respective class URI from the SRO ontology as the annotation feature. You can find the URLs of the gold standard documents using the file.
Note that we could not provide the annotated documents of the SePublica corpus as we have to clear the rights to the documents with the individual authors first. Instead, the SePublica_GS.csv file in the gold standard directory provides the annotations and their type in a comma-separated format that can be mapped to the original documents using the provided start and end offsets in text.
1.3. Knowledge Base
You can download the complete (zip-compressed) knowledge base as we generated it for the paper, in N-Quads format (1,086,051 triples). It was generated with Jena's tdbdump command, but should load fine into other, non-Jena triplestores as well. To load it into a new KB, create an empty directory (e.g., /tmp/tdb) and issue:
- tdbloader --loc=/tmp/tdb triples.nq
For more information on tdbloader, please refer to Apache Jena's TDB Command-line Utilities page.
2. Software
All software described in the paper is available under open source licenses.
2.1. Required Third-Party Software
- You need a JDK (Java 7 or better), as well as an installation of Apache Ant.
- You also need Apache Jena, version 2.13 or better.
- If you want to run the text mining pipeline, you must have GATE, version 8.1 or better, installed. See the GATE homepage for instructions on how to install and run GATE.
- Also, for the text mining pipeline, you need access to a DBpedia Spotlight installation (RESTful interface). Since the output highly depends on the model used for Spotlight, if you want to reproduce our results, you will need to use the exact same model as in our paper: en_2+2.
- For generating the result data table as shown in the paper using the provided stats-reporter tool, you also need a standard LaTeX distribution installed.
2.2. Mapping file
The lodexporter-mapping.zip file contains the rules for mapping GATE annotations to RDF triples. You will need to import them into your TDB-based triplestore, e.g., using the tdbloader command. E.g., if you want to store your KB into /tmp/tdb, use the command:
- tdbloader --loc=/tmp/tdb lodexporter-mapping.rdf
2.3. Text Mining Pipeline
The text mining pipeline described in the paper is provided as a ZIP file. Unzip the downloaded package on your workstation and follow the instructions below to reproduce our experiments: (Note: the pipeline has been tested on Linux and MacOS X):
- You must have created a TDB-based triplestore and loaded our mapping rules as described above.
- Start GATE (v8.1 or better). Choose File → Restore Application from File. Then browse to where you unzipped the pipeline and choose the provided XGAPP file.
- Once the pipeline is loaded, you can open any document (e.g., the gold corpora provided here).
- Create a new corpus and add the documents.
- Double-click on the PeerJ_CompSci2015_Semantic_Lit_Mining pipeline under Applications. Choose the new corpus you created above from the dropdown and click Run this Application button.
- Once the pipeline is finished you can open the documents and examine their annotations.
- In order to examine the generated triples, first close the GATE application. Then you can either check the triples using Jena's tdbdump command or publish it through a Fuseki server.
2.4. Table generation
Nobody likes writing (and updating) tables by hand, so we use a basic Java program to (i) query the generated KB and (ii) format the results into a LaTeX table. You can easily reproduce the table shown in the paper by creating a TDB-based triplestore that contains the triples provided above and running the included stats-reporter Java program by calling ant. Note: by default the program looks for the triplestore in /tmp/tdb. You can change this in the build.properties file.
After calling ant, the program will compile, run, and fill the table cells through the SPARQL queries contained in the source code. You can find the resulting table in the output directory (default, change it in build.properties) in the file corpus_table.tex. After compiling the provided StatsReporter.tex main file including this table (e.g., with pdflatex), you should see an output like this:
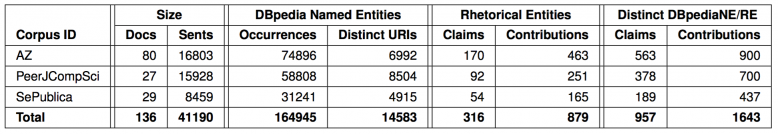 Table as generated for our PeerJ CompiSci 2015 paper using the stats-reporter tool
Table as generated for our PeerJ CompiSci 2015 paper using the stats-reporter tool
3. Knowledge Base Queries
In the "Application" section of our paper, there are three scenarios, where the user interacts with the populated knowledge base in order to retrieve documents related to her task at hand. In this page, you can find each query and execute it yourself by copying and pasting it to the query box below.
As a running example, let us imagine a use case: A user wants to write a literature review from a given set of documents about a specific topic. Ordinarily, the user has to read all of the retrieved documents in order to evaluate their relevance to her task — a cumbersome and time-consuming task. However, using our approach the user can directly query for the rhetorical type that she needs from the system. We demonstrate this with the example queries shown below.
Scenario 1: The user has downloaded a set of new articles from the web. Before reading each article thoroughly, she would like to obtain a summary of the contributions of all articles, so she can decide which articles are relevant to her task.
- PREFIX pubo: <http://lod.semanticsoftware.info/pubo/pubo#>
- PREFIX sro: <http://salt.semanticauthoring.org/ontologies/sro#>
- PREFIX rdf: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
- PREFIX cnt: <http://www.w3.org/2011/content#>
- SELECT ?paper ?content WHERE {
- ?paper pubo:hasAnnotation ?rhetoricalEntity .
- ?rhetoricalEntity rdf:type sro:Contribution .
- ?rhetoricalEntity cnt:chars ?content }
Scenario 2: From the set of downloaded articles, the user would like to find only those articles that have a contribution mentioning `linked data'.
- PREFIX pubo: <http://lod.semanticsoftware.info/pubo/pubo#>
- PREFIX sro: <http://salt.semanticauthoring.org/ontologies/sro#>
- PREFIX rdf: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
- PREFIX rdfs: <http://www.w3.org/2000/01/rdf-schema#>
- PREFIX cnt: <http://www.w3.org/2011/content#>
- PREFIX dbpedia: <http://dbpedia.org/resource/>
- SELECT DISTINCT ?paper ?content WHERE {
- ?paper pubo:hasAnnotation ?rhetoricalEntity .
- ?rhetoricalEntity rdf:type sro:Contribution .
- ?rhetoricalEntity pubo:containsNE ?ne.
- ?ne rdfs:isDefinedBy dbpedia:Linked_data .
- ?rhetoricalEntity cnt:chars ?content } ORDER BY ?paper
Scenario 3: The user would like to find only those articles that have a contribution mentioning topics related to `linked data'.
(Please note that this query uses the public DBpedia endpoint and thus may take longer to execute, or fail if the endpoint is offline.)
- PREFIX pubo: <http://lod.semanticsoftware.info/pubo/pubo#>
- PREFIX sro: <http://salt.semanticauthoring.org/ontologies/sro#>
- PREFIX rdf: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
- PREFIX rdfs: <http://www.w3.org/2000/01/rdf-schema#>
- PREFIX cnt: <http://www.w3.org/2011/content#>
- PREFIX dcterms: <http://purl.org/dc/terms/>
- PREFIX dbpedia: <http://dbpedia.org/resource/>
- SELECT ?paper ?content WHERE {
- SERVICE <http://dbpedia.org/sparql> {
- dbpedia:Linked_data dcterms:subject ?category .
- ?subject dcterms:subject ?category . }
- ?paper pubo:hasAnnotation ?rhetoricalEntity .
- ?rhetoricalEntity rdf:type sro:Contribution .
- ?rhetoricalEntity pubo:containsNE ?ne.
- ?ne rdfs:isDefinedBy ?subject .
- ?rhetoricalEntity cnt:chars ?content }
References
| Attachment | Size |
|---|---|
| gate-pipeline.zip | 1.21 MB |
| corpus-index-20151109.txt | 3.53 KB |
| lodexporter-mapping.zip | 1.47 KB |
| knowledge-base-20151109.zip | 7.61 MB |
| goldstandard-corpus-index-20151109.txt | 2.88 KB |
| goldstandard-20151109.zip | 1.08 MB |
| stats-reporter.zip | 25.79 KB |



